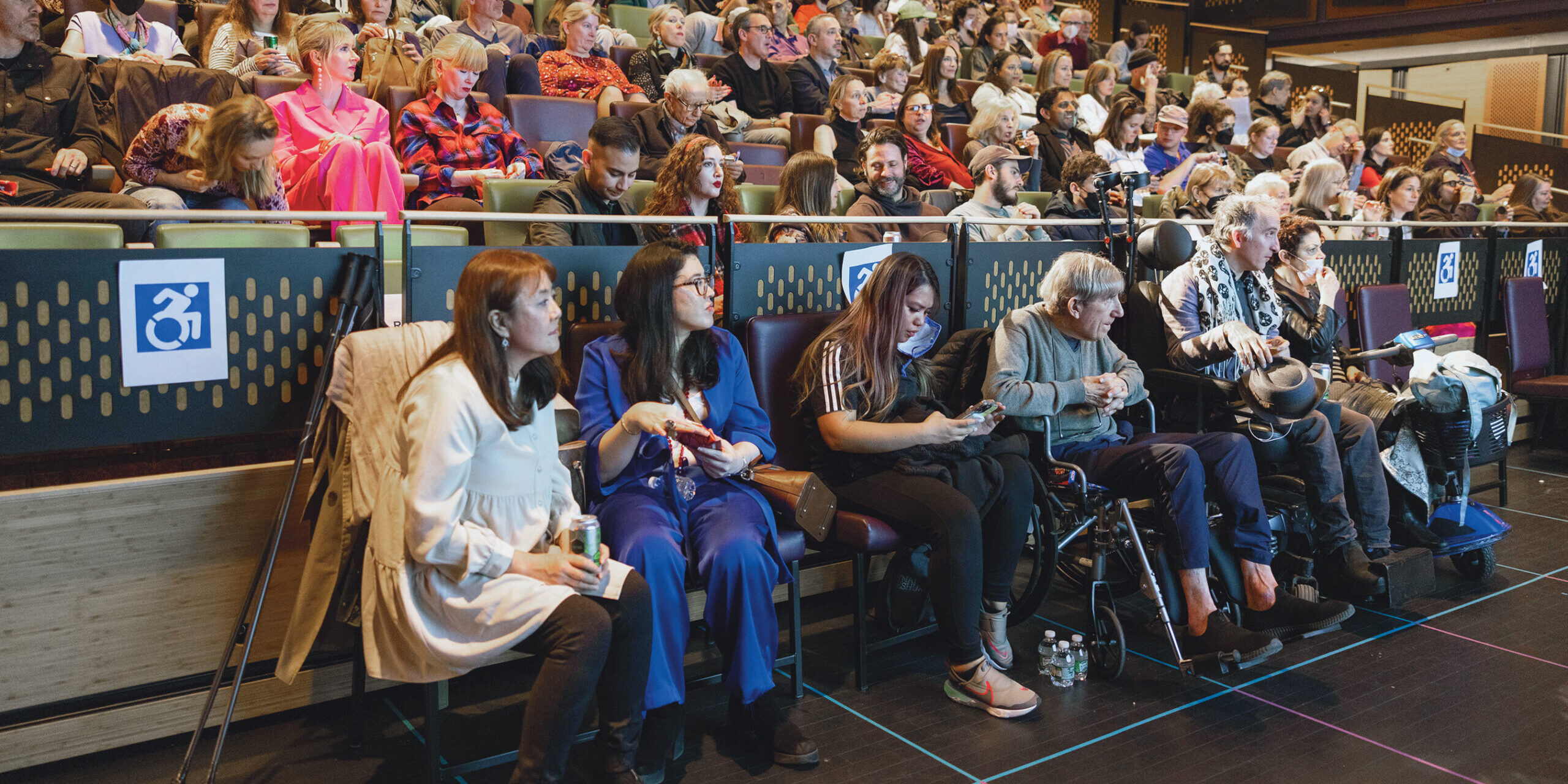
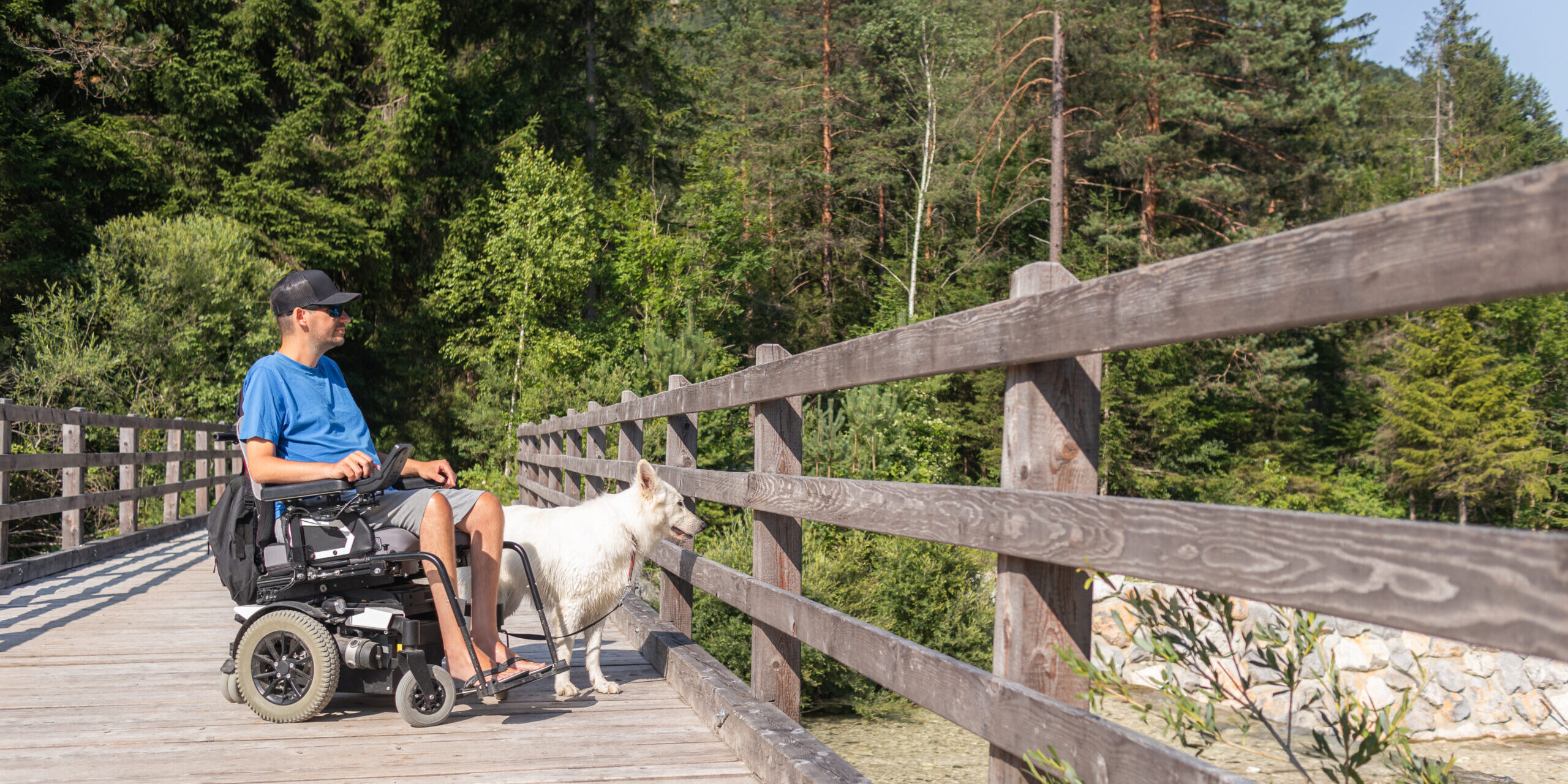
Quest Media is an innovative adaptive lifestyle platform from MDA. With the power of this platform, we foster awareness and empowerment and have important conversations with experts, thought leaders, and members of the neuromuscular disease community about topics that matter to them and to the larger community of individuals with disabilities.
MUST-READ FEATURES
QUEST PODCAST
The Quest podcast, proudly presented by the Muscular Dystrophy Association, is part of the Quest family of content. Hosted by Quest Editor-in-Chief, motivational speaker and writer Mindy Henderson.
Episode 40- Unlocking Access and Inspiring Action with Sophie Morgan
In this Quest Podcast episode, we chat with world-renowned advocate, entrepreneur, TV personality and producer, Sophie Morgan. She devotes her time and expertise to create inclusive spaces for those with disabilities and deliver advice, inspire action, and make us feel closer together while sharing stories of resilience and positivity. Sophie is a co-founder of Making…
Episode 39- Behind the Scenes: A Look at the Science and Research for New Treatments
In this Quest Podcast episode, we chat with geneticist, Dr. Jeffrey Chamberlain and the Chief Research Officer of the Muscular Dystrophy Association, Dr. Sharon Hesterlee. Both have devoted their time and expertise to create and move forward research and treatments for neuromuscular diseases. Their goal is to create successful treatments and eventually a cure for…
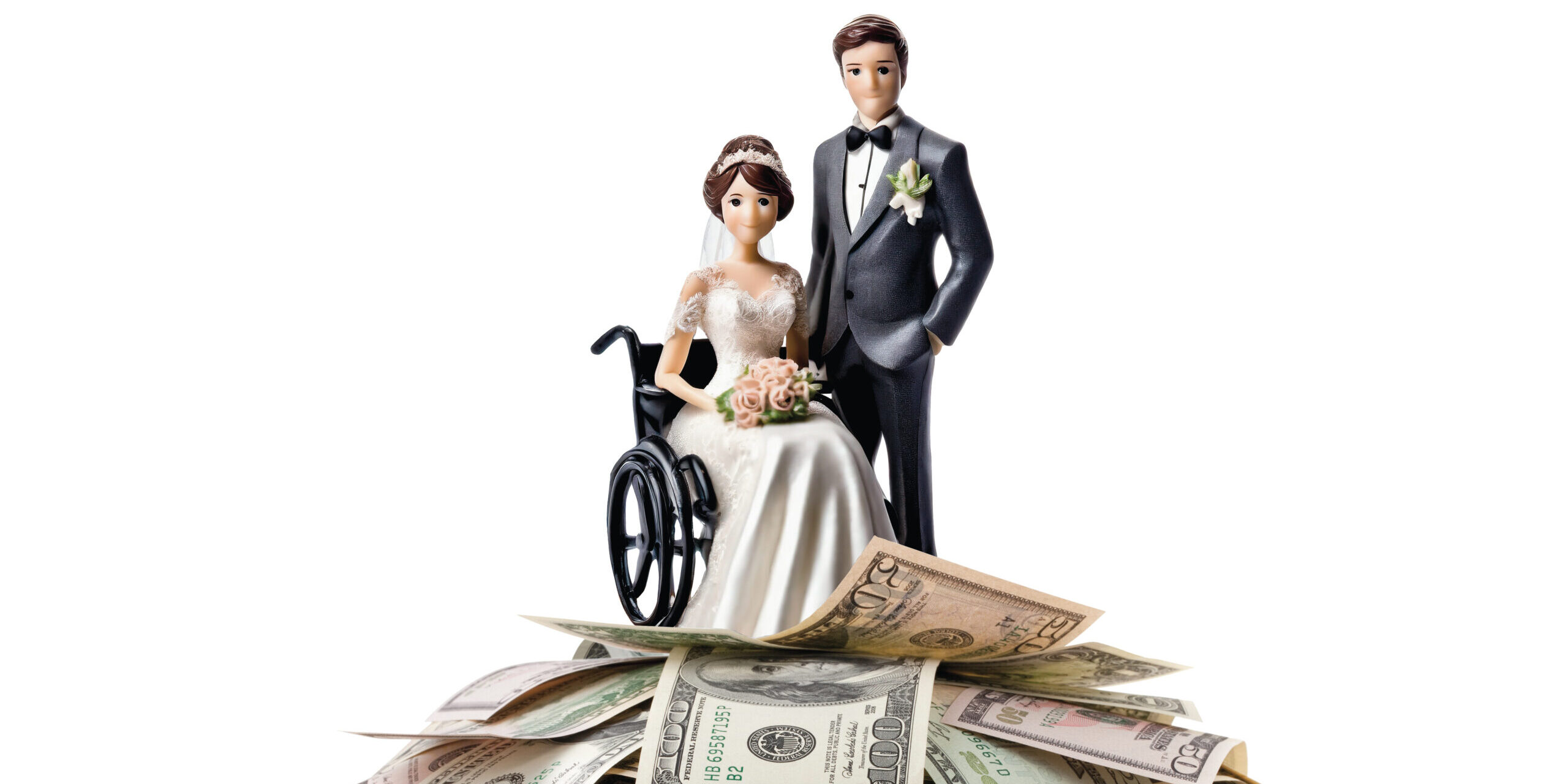
Episode 38: Love Made Simple with DateAbility’s Alexa and Jacqueline Child
In this Quest Podcast episode, we chat with the founders of DateAbility, a dating application geared towards individuals with disabilities and chronic illnesses. Alexa and Jacqueline Child have devoted their time to create a safe and accepting space that allows individuals to create meaningful connections. Their goal is to make love accessible for everyone. These…